Before diving into the various techniques of model validation, we need to understand why validating is important. Through validating we can find out how well our model works by comparing the predicted value and the actual value and all this is done with the training dataset. but the problem is that the training dataset can easily overfit the model (giving fake accurate results when performing model evaluation) and thus will not give good results with unseen data.

Thus , as a solution we divide the original data in such a way that we train the model on the "training set" such that it perform reasonably well in the model evaluation but also give good result when the evaluation is done with the other part of original dataset known as testing dataset or validation dataset. In this way we can find out when our model is overfitting and hence we can build a model which will work reasonably well on the unseen data.
Overfitting is the scenario where the predictor becomes too complex and fits the data too well, causing the model to overfit. This is usually due to the noise and other patterns present in the dataset which is not the case with the general data and hence the model fails miserably with unseen data. And thus, we require a function that is not very accurate in the training phase but is adequately accurate in the testing phase. Similarly, underfitting is the problem when the function is not complex enough and fails to generalize the pattern, thus giving inaccurate results in both training and testing phase. Thus we need to strike a balance between the complexity of the function. and this is where cross validation comes into picture.
In simple terms, Cross-Validation is a technique used to assess how well our Machine learning models perform on unseen data. Cross validation is a model evaluation method that is better than residuals. The problem with residual evaluations is that they do not give an indication of how well the learner will do when it is asked to make new predictions for data it has not already seen. One way to overcome this problem is to not use the entire data set when training a learner. Some of the data is removed before training begins. Then when training is done, the data that was removed can be used to test the performance of the learned model on ``new'' data. This is the basic idea for a whole class of model evaluation methods called cross validation.
The two most commonly used methods for cross-validation are the holdout method and k-fold cross-validation.
The holdout method is the simplest kind of cross validation. The data set is separated into two sets, called the training set and the testing set. The function approximator fits a function using the training set only. Then the function approximator is asked to predict the output values for the data in the testing set (it has never seen these output values before). The errors it makes are accumulated as before to give the mean absolute test set error, which is used to evaluate the model. The advantage of this method is that it is usually preferable to the residual method and takes no longer to compute. However, its evaluation can have a high variance. The evaluation may depend heavily on which data points end up in the training set and which end up in the test set, and thus the evaluation may be significantly different depending on how the division is made. The steps in this method are:

The data can be divided into 70-30 or 60-40, 75-25 or 80-20, or even 50-50 depending on the use case. As a rule, the proportion of training data has to be larger than the test data.
Implementation in Python:
We can now quickly sample a training set while holding out 40% of the data for testing (evaluating) our classifier using train_test_split from sklearn.model_selection library.
import numpy as np from sklearn.model_selection import train_test_split from sklearn import datasets from sklearn import svm
X, y = datasets.load_iris(return_X_y=True) print(X.shape, y.shape)
((150, 4), (150,))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=0) print(X_train.shape, y_train.shape) print(X_test.shape, y_test.shape)
((90, 4), (90,)) ((60, 4), (60,))
clf = svm.SVC(kernel='linear', C=1).fit(X_train, y_train) print(clf.score(X_test, y_test))
0.96...
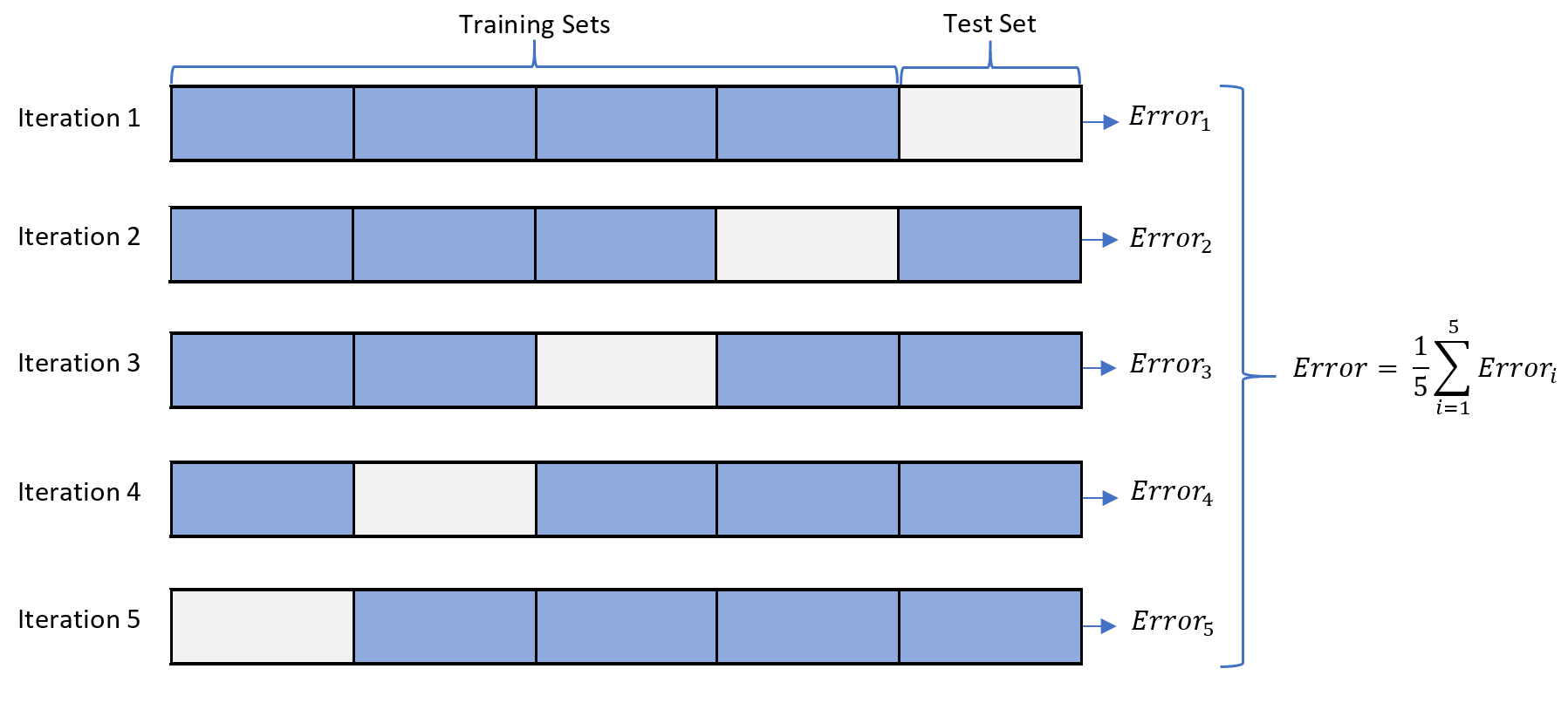
K-fold cross validation is one way to improve over the holdout method. The data set is divided into k subsets, and the holdout method is repeated k times. Each time, one of the k subsets is used as the test set and the other k-1 subsets are put together to form a training set. Then the average error across all k trials is computed. The advantage of this method is that it matters less how the data gets divided. Every data point gets to be in a test set exactly once, and gets to be in a training set k-1 times. The variance of the resulting estimate is reduced as k is increased. The disadvantage of this method is that the training algorithm has to be rerun from scratch k times, which means it takes k times as much computation to make an evaluation. A variant of this method is to randomly divide the data into a test and training set k different times. The advantage of doing this is that you can independently choose how large each test set is and how many trials you average over.
Implementation in Python:
#Importing required libraries
from sklearn.datasets import load_breast_cancer
import pandas as pd
from sklearn.model_selection import KFold
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
#Loading the dataset
data = load_breast_cancer(as_frame = True)
df = data.frame
X = df.iloc[:,:-1]
y = df.iloc[:,-1]
#Implementing cross validation
k = 5
kf = KFold(n_splits=k, random_state=None)
model = LogisticRegression(solver= 'liblinear')
acc_score = []
for train_index , test_index in kf.split(X):
X_train , X_test = X.iloc[train_index,:],X.iloc[test_index,:]
y_train , y_test = y[train_index] , y[test_index]
model.fit(X_train,y_train)
pred_values = model.predict(X_test)
acc = accuracy_score(pred_values , y_test)
acc_score.append(acc)
avg_acc_score = sum(acc_score)/k
print('accuracy of each fold - {}'.format(acc_score))
print('Avg accuracy : {}'.format(avg_acc_score))
accuracy of each fold - [0.9122807017543859, 0.9473684210526315, 0.9736842105263158, 0.9736842105263158, 0.9557522123893806] Avg accuracy : 0.952553951249806
Leave-one-out cross validation is K-fold cross validation taken to its logical extreme, with K equal to N, the number of data points in the set. That means that N separate times, the function approximator is trained on all the data except for one point and a prediction is made for that point. As before the average error is computed and used to evaluate the model. The evaluation given by leave-one-out cross validation error (LOO-XVE) is good, but at first pass it seems very expensive to compute. Fortunately, locally weighted learners can make LOO predictions just as easily as they make regular predictions. That means computing the LOO-XVE takes no more time than computing the residual error and it is a much better way to evaluate models. We will see shortly that Vizier relies heavily on LOO-XVE to choose its metacodes.
implementation in python:
import numpy as np from sklearn.model_selection import LeaveOneOut X = np.array([[1, 2], [3, 4]]) y = np.array([1, 2]) loo = LeaveOneOut() print(loo.get_n_splits(X))
2
for train_index, test_index in loo.split(X):
print("TRAIN:", train_index, "TEST:", test_index)
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
print(X_train, X_test, y_train, y_test)
TRAIN: [1] TEST: [0] [[3 4]] [[1 2]] [2] [1] TRAIN: [0] TEST: [1] [[1 2]] [[3 4]] [1] [2]
Before diving deep into stratified cross-validation, it is important to know about stratified sampling. Stratified sampling is a sampling technique where the samples are selected in the same proportion (by dividing the population into groups called ‘strata’ based on a characteristic) as they appear in the population. Implementing the concept of stratified sampling in cross-validation ensures the training and test sets have the same proportion of the feature of interest as in the original dataset. Doing this with the target variable ensures that the cross-validation result is a close approximation of generalization error.
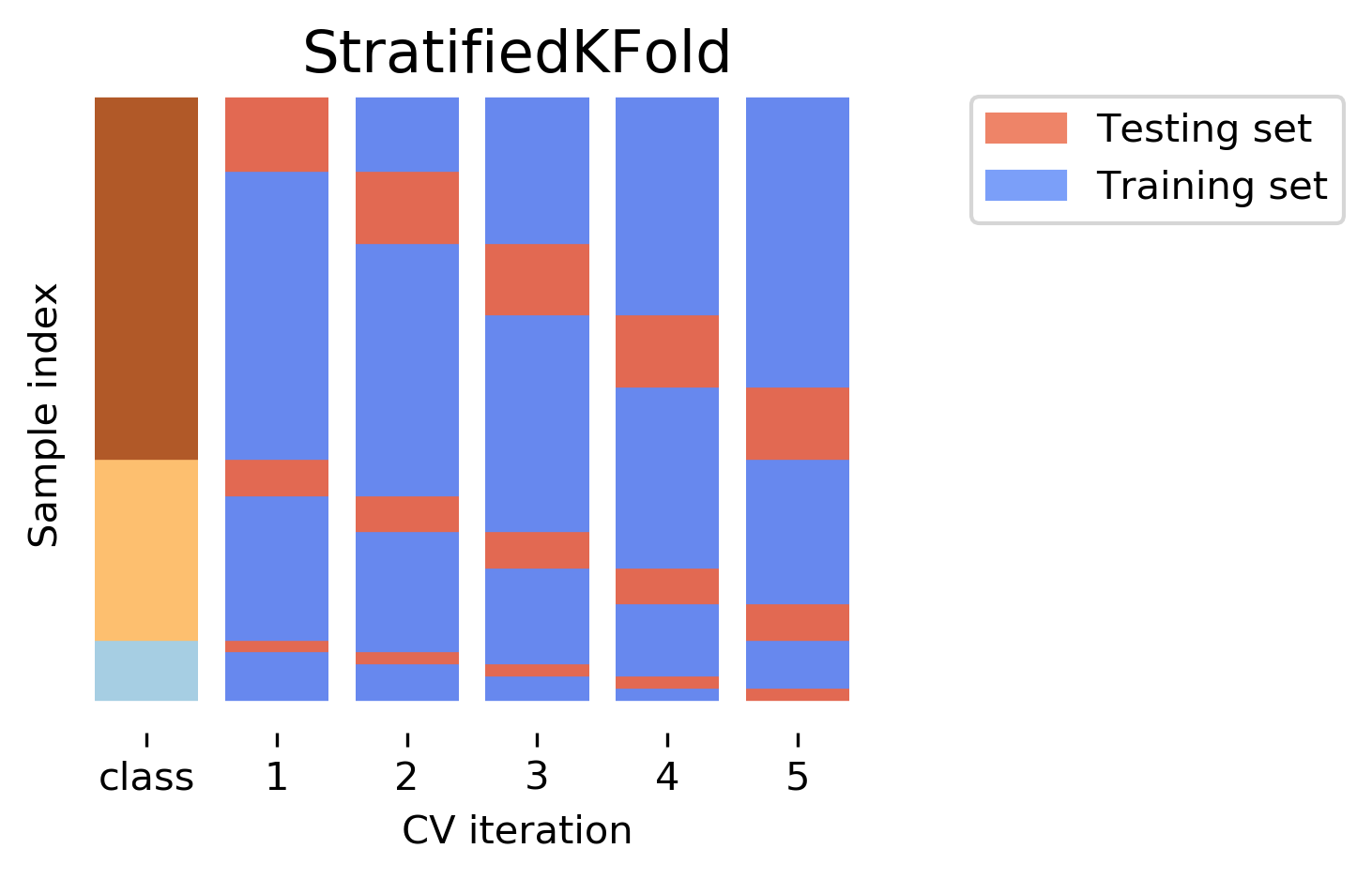
implementation in python:
import pandas as pd
from sklearn.model_selection import StratifiedKFold
from sklearn.linear_model import LogisticRegression
dataset = pd.read_csv('/content/diabetes.csv')
skf = StratifiedKFold(n_splits=10)
model = LogisticRegression(solver='newton-cg')
x = dataset
y = dataset.Outcome
def training(train, test, fold_no):
x_train = train.drop(['Outcome'],axis=1)
y_train = train.Outcome
x_test = test.drop(['Outcome'],axis=1)
y_test = test.Outcome
model.fit(x_train, y_train)
score = model.score(x_test,y_test)
print('For Fold {} the accuracy is {}'.format(str(fold_no),score))
fold_no = 1
for train_index,test_index in skf.split(x, y):
train = dataset.iloc[train_index,:]
test = dataset.iloc[test_index,:]
training(train, test, fold_no)
fold_no += 1

When we consider the test error rate estimates, K-Fold Cross Validation gives more accurate estimates than Leave One Out Cross-Validation. Whereas Hold One Out CV method usually leads to overestimates of the test error rate, because in this approach, only a portion of the data is used to train the machine learning model.
When it comes to bias, the Leave One Out Method gives unbiased estimates because each training set contains n-1 observations (which is pretty much all of the data). K-Fold CV leads to an intermediate level of bias depending on the number of k-folds when compared to LOOCV but it’s much lower when compared to the Hold Out Method.
To conclude, the Cross-Validation technique that we choose highly depends on the use case and bias-variance trade-off.